maint_debian-med package set for unstable/amd64
Debian package sets:
desktop package sets:
Debian distribution package sets:
maintenance team package sets:
-
 maint_debian-accessibility
maint_debian-accessibility
-
 maint_debian-boot
maint_debian-boot
-
 maint_debian-lua
maint_debian-lua
-
 maint_debian-med
maint_debian-med
-
 maint_debian-ocaml
maint_debian-ocaml
-
 maint_debian-on-mobile-maintainers
maint_debian-on-mobile-maintainers
-
 maint_debian-python
maint_debian-python
-
 maint_debian-qa
maint_debian-qa
-
 maint_debian-science
maint_debian-science
-
 maint_debian-x
maint_debian-x
-
 maint_pkg-android-tools-devel
maint_pkg-android-tools-devel
-
 maint_pkg-erlang-devel
maint_pkg-erlang-devel
-
 maint_pkg-fonts-devel
maint_pkg-fonts-devel
-
 maint_pkg-games-devel
maint_pkg-games-devel
-
 maint_pkg-golang-maintainers
maint_pkg-golang-maintainers
-
 maint_pkg-grass-devel
maint_pkg-grass-devel
-
 maint_pkg-haskell-maintainers
maint_pkg-haskell-maintainers
-
 maint_pkg-java-maintainers
maint_pkg-java-maintainers
-
 maint_pkg-javascript-devel
maint_pkg-javascript-devel
-
 maint_pkg-multimedia-maintainers
maint_pkg-multimedia-maintainers
-
 maint_pkg-perl-maintainers
maint_pkg-perl-maintainers
-
 maint_pkg-php-pear
maint_pkg-php-pear
-
 maint_pkg-openstack
maint_pkg-openstack
-
 maint_pkg-r
maint_pkg-r
-
 maint_pkg-ruby-extras-maintainers
maint_pkg-ruby-extras-maintainers
-
 maint_pkg-rust-maintainers
maint_pkg-rust-maintainers
-
 maint_reproducible-builds
maint_reproducible-builds
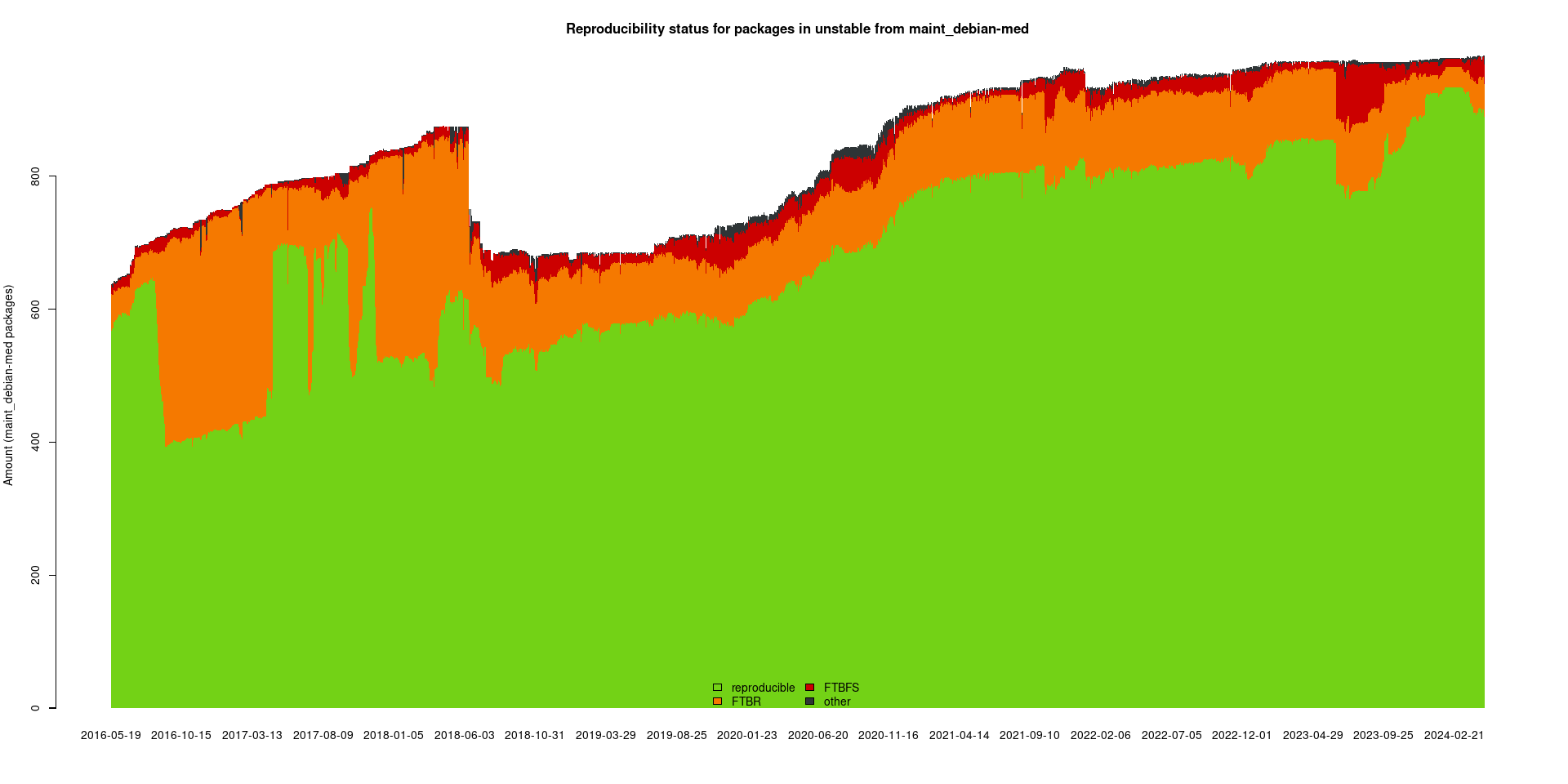
The package set maint_debian-med in
unstable/amd64 consists of 982 packages:
 44 (4.5%) packages
failed to build reproducibly:
libcifpp
filtlong
pydicom
mrtrix3
liblemon
shapeit4
twopaco
btllib
jebl2
librostlab
tree-puzzle
bmtk
heudiconv
seqan3
unicycler
fis-gtm
cwltool
nitime
dipy
emboss
dcmtk
gbrowse
hmmer
igraph
metastudent-data
macsyfinder
odil
treeview
porechop
iqtree
dicom3tools
gdcm
python-pysam
mapsembler2
ball
libpll
salmon
seqan2
ncbi-blast+
segemehl
kleborate
ncbi-igblast
parallel
gatb-core
44 (4.5%) packages
failed to build reproducibly:
libcifpp
filtlong
pydicom
mrtrix3
liblemon
shapeit4
twopaco
btllib
jebl2
librostlab
tree-puzzle
bmtk
heudiconv
seqan3
unicycler
fis-gtm
cwltool
nitime
dipy
emboss
dcmtk
gbrowse
hmmer
igraph
metastudent-data
macsyfinder
odil
treeview
porechop
iqtree
dicom3tools
gdcm
python-pysam
mapsembler2
ball
libpll
salmon
seqan2
ncbi-blast+
segemehl
kleborate
ncbi-igblast
parallel
gatb-core
 18 (1.8%) packages
failed to build from source:
pyranges
beast2-mcmc
vg
q2-feature-classifier
libsbml
acedb
cmtk
mlv-smile
pynn
intake
toil
python-array-api-compat
pplacer
unifrac-tools
mia
freebayes
anfo
consensuscore
18 (1.8%) packages
failed to build from source:
pyranges
beast2-mcmc
vg
q2-feature-classifier
libsbml
acedb
cmtk
mlv-smile
pynn
intake
toil
python-array-api-compat
pplacer
unifrac-tools
mia
freebayes
anfo
consensuscore



 8 (0.8%) packages
are either in depwait state, blacklisted, not for us, or cannot be downloaded:
psychopy
python-cogent
umap-learn
python-loompy
pysurfer
pymia
mrc
sourmash
8 (0.8%) packages
are either in depwait state, blacklisted, not for us, or cannot be downloaded:
psychopy
python-cogent
umap-learn
python-loompy
pysurfer
pymia
mrc
sourmash
 912 (92.9%) packages
successfully build reproducibly:
abacas
abpoa
abyss
adapterremoval
adun.app
aegean
aeonbits-owner
aevol
aghermann
alien-hunter
allelecount
alter-sequence-alignment
altree
amap-align
amide
ampliconnoise
andi
ants
any2fasta
aragorn
arden
argh
argtable2
ariba
artemis
artfastqgenerator
art-nextgen-simulation-tools
assembly-stats
assemblytics
ataqv
atropos
augur
augustus
autodocksuite
autodock-vina
axe-demultiplexer
baitfisher
bali-phy
bambamc
bamclipper
bamkit
bamtools
bandage
barrnap
bart
bart-view
bbhash
bbmap
bcalm
bcftools
beads
beagle
beast-mcmc
bedops
bedtools
berkeley-express
bifrost
bioawk
biobambam2
bio-eagle
biojava4-live
biojava6-live
biojava-live
biomaj3
biomaj3-cli
biomaj3-core
biomaj3-daemon
biomaj3-download
biomaj3-process
biomaj3-user
biomaj3-zipkin
bioperl
bioperl-run
bio-rainbow
biosig
biosquid
biosyntax
bio-tradis
bio-vcf
bitseq
blasr
bolt-lmm
bowtie
bowtie2
boxshade
bppphyview
bppsuite
brian
brig
busco
bustools
bwa
camitk
camp
canu
capsule-maven-nextflow
capsule-nextflow
cassiopee
castxml
cat-bat
cct
cdbfasta
cde
cd-hit
centrifuge
cgview
changeo
charls
chip-seq
chromhmm
chromimpute
ciftilib
cif-tools
circlator
circos
circos-tools
civetweb
clearcut
clonalframe
clonalframeml
clonalorigin
clustalo
clustalw
clustalx
cnvkit
codonw
coils
concavity
concurrentqueue
conda-package-handling
conda-package-streaming
conservation-code
covtobed
crac
ctdconverter
ctdopts
ctffind
ctn
ctsim
cutesv
cwlformat
cwltest
cycle
cyvcf2
daligner
damapper
dascrubber
dawg
dazzdb
dcm2niix
dcmstack
debian-med
deblur
delly
density-fitness
dextractor
dialign
dialign-t
diamond-aligner
dicomscope
discosnp
disulfinder
dnaclust
dnapi
dnarrange
drmaa
drop-seq
dssp
dwgsim
ea-utils
ecopcr
edfbrowser
edflib
edtsurf
eegdev
eigensoft
elastix
elph
embassy-domainatrix
embassy-domalign
embassy-domsearch
emboss-explorer
e-mem
emmax
emperor
epcr
estscan
examl
exonerate
fast5
fasta3
fastani
fastaq
fastdnaml
fastlink
fastml
fastp
fastqc
fastq-pair
fastqtl
fasttree
fermi-lite
ffindex
figtree
fitgcp
flash
flexbar
flye
freecontact
fsa
fsm-lite
g2
galileo
garli
gasic
gatk-bwamem
gatk-fermilite
gdpc
gemma
genometester
genomethreader
genometools
genomicsdb
gentle
getdata
gfapy
gff2aplot
gff2ps
gffread
ggd-utils
ghmm
gifticlib
gjh-asl-json
glam2
gmap
gnumed-client
gnumed-server
grabix
graphlan
grinder
gsort
gubbins
gwama
gwyddion
harmonypy
harvest-tools
hdmf
hhsuite
hilive
hinge
hisat2
hmmer2
hnswlib
hopscotch-map
htscodecs
htseq
htsjdk
htslib
hts-nim-tools
hunspell-en-med
hyphy
icb-utils
idba
idseq-bench
igor
igv
iitii
illustrate
imagej
imbalanced-learn
indelible
infernal
insighttoolkit5
insilicoseq
invesalius
ipig
ismrmrd#
itkadaptivedenoising
itkgenericlabelinterpolator
iva
ivar
jaligner
jam-lib
jellyfish
jellyfish1
jheatchart
jmodeltest
kalign
kallisto
kaptive
kineticstools
king
king-probe
kissplice
klustakwik
kma
kmc
kmer
kmerresistance
kraken
kraken2
lagan
lamarc
lamassemble
lambda-align
lambda-align2
last-align
lastz
lefse
libace-perl
libamplsolver
libargparse
libargs
libatomicbitvector
libatomic-queue
libbigwig
libbio-alignio-stockholm-perl
libbio-cluster-perl
libbio-coordinate-perl
libbio-das-lite-perl
libbio-db-biofetch-perl
libbio-db-embl-perl
libbio-db-hts-perl
libbio-db-ncbihelper-perl
libbio-db-seqfeature-perl
libbio-featureio-perl
libbio-graphics-perl
libbio-mage-perl
libbio-mage-utils-perl
libbioparser-dev
libbiosoup-dev
libbio-tools-run-remoteblast-perl
libbio-variation-perl
libbitarray
libbpp-core
libbpp-phyl
libbpp-phyl-omics
libbpp-popgen
libbpp-qt
libbpp-raa
libbpp-seq
libbpp-seq-omics
libcereal
libchado-perl
libcolt-free-java
libctapimkt
libdisorder
libdistlib-java
libdivsufsort
libedlib
libfastahack
libflathashmap
libfreecontact-perl
libgclib
libgdf
libgenome
libgenome-model-tools-music-perl
libgenome-perl
libgff
libgkarrays
libgoby-java
libgo-perl
libgtextutils
libgzstream
libhac-java
libhat-trie
libhmsbeagle
libhpptools
libhttp-nio-java
libics
libips4o
libjloda-java
libjung-free-java
libkmlframework-java
libla4j-java
libleidenalg
libmaus2
libmcfp
libmems
libmialm
libminc
libmjson-java
libmmap-allocator
libmmmulti
libmurmurhash
libmuscle
libncl
libnewuoa
libomp-jonathonl
liboptions-java
libpal-java
libpdb-redo
libpj-java
libpsortb
libqes
librandom123
librcsb-core-wrapper
librdp-taxonomy-tree-java
librg-blast-parser-perl
librg-exception-perl
librg-utils-perl
librostlab-blast
libsecrecy
libseqlib
libshrinkwrap
libsis-base-java
libslow5lib
libsmithwaterman
libsort-key-top-perl
libssw
libstatgen
libstreamvbyte
libtabixpp
libtecla
libtfbs-perl
libthread-pool
libvbz-hdf-plugin
libvcflib
libvistaio
libwfa2
libxdf
libzeep
libzerg
libzerg-perl
lighter
loki
ltrsift
lucy
lumpy-sv
macs
maffilter
mafft
malt
mapdamage
maq
maqview
mash
maude
mauve-aligner
maxflow
mcaller
mcl
mcl14
mecat2
medicalterms
megadepth
megahit
megan-ce
melting
mencal
metabat
metaeuk
metaphlan
metaphlan2-data
metastudent
metastudent-data-2
mhap
mialmpick
microbegps
microbiomeutil
milib
minc-tools
mindthegap
minia
miniasm
minimac4
minimap
minimap2
mipe
mira
mirtop
mmlib
mmseqs2
mosdepth
mothur
mptp
mrbayes
mricron
multiqc
mummer
murasaki
muscle
muscle3
mustang
nanofilt
nanolyse
nanook
nanopolish
nanostat
nanosv
ncbi-acc-download
ncbi-entrez-direct
ncbi-seg
ncbi-tools6
ncbi-vdb
neo
neobio
ngmlr
nibabel
nifticlib
nim-hts
nim-kexpr
nim-lapper
nipy
nipype
njplot
norsnet
norsp
ntcard
nthash
nutsqlite
nxtrim
obitools
odin
ont-fast5-api
opencfu
openslide
openslide-python
optimir
orthanc
orthanc-dicomweb
orthanc-gdcm
orthanc-imagej
orthanc-mysql
orthanc-neuro
orthanc-postgresql
orthanc-python
orthanc-webviewer
orthanc-wsi
oscar
pairtools
pal2nal
paleomix
paml
papyrus
paraclu
parafly
parallel-fastq-dump
parasail
parsinsert
parsnp
paryfor
patman
patsy
pbbam
pbcopper
pbdagcon
pbseqlib
pbsim
pbsuite
pcalendar
pdb2pqr
peptidebuilder
perlprimer
perm
pftools
phast
phipack
phybin
phylip
phylonium
phyml
physamp
phyutility
phyx
picard-tools
picopore
pigx-rnaseq
piler
pilercr
pilon
pinfish
pique
pirs
pixelmed
pixelmed-codec
pizzly
placnet
plasmidid
plasmidomics
plasmidseeker
plast
plastimatch
plink
plink1.9
plink2
plip
poa
populations
poretools
praat
prank
predictnls
presto
prime-phylo
primer3
prinseq-lite
proalign
probabel
probalign
probcons
proda
prodigal
profbval
profisis
profnet
profphd-utils
proftmb
progressivemauve
prokka
propka
proteinortho
prottest
provean
pscan-chip
pscan-tfbs
psignifit
psortb
pullseq
pvrg-jpeg
pybel
pybigwig
pychopper
pycoqc
pycorrfit
pyensembl
pyfastx
pynwb
pyode
pyqi
pyscanfcs
python-alignlib
python-anndata
python-awkward
python-bcbio-gff
python-bel-resources
python-bids-validator
python-bioblend
python-bioframe
python-biom-format
python-biopython
python-bioregistry
python-biotools
python-bx
python-cgecore
python-cgelib
python-cigar
python-cobra
python-cooler
python-csb
python-curies
python-cutadapt
python-cykhash
python-datacache
python-deeptools
python-deeptoolsintervals
python-dendropy
python-dicompylercore
python-dnaio
python-epimodels
python-ete3
python-etelemetry
python-fitbit
python-freecontact
python-geneimpacts
python-gffutils
python-gtfparse
python-hl7
python-hmmlearn
python-intervaltree-bio
python-iow
python-leidenalg
python-mne
python-nanoget
python-nanomath
python-pairix
python-pangolearn
python-parasail
python-pauvre
python-pbcommand
python-pbcore+
python-prefixed
python-py2bit
python-pyani
python-pybedtools
python-pycosat
python-pyfaidx
python-pymummer
python-pyspoa
python-pyvcf
python-questplus
python-ruffus
python-scitrack
python-screed
python-skbio
python-sqt
python-tinyalign
python-treetime
python-trx-python
python-wdlparse
pyxid
pyxnat
q2-alignment
q2cli
q2-cutadapt
q2-dada2
q2-demux
q2-diversity-lib
q2-emperor
q2-feature-table
q2-fragment-insertion
q2-metadata
q2-phylogeny
q2-quality-control
q2-quality-filter
q2-sample-classifier
q2-taxa
q2templates
q2-types
qcat
qcumber
qiime
qrisk2
qsopt-ex
qtltools
quicktree
quitcount
quorum
racon
radiant
ragout
rambo-k
rampler
rapmap
raster3d
rate4site
raxml
ray
rdp-alignment
rdp-classifier
rdp-readseq
readerwriterqueue
readseq
readucks
reapr
recan
relacy
relion
repeatmasker-recon
reprof
resfinder
resfinder-db
rnahybrid
rna-star
roary
rockhopper
roguenarok
routine-update
rsem
rtax
ruby-rgfa
runcircos-gui
saint
salmid
sambamba
samblaster
samclip
samtools
samtools-legacy
savvy
sbmltoolbox
scoary
scrappie
scrm
scythe
seaview
seer
seirsplus
sepp
seqan-needle
seqan-raptor
seqkit
seqmagick
seqprep
seqsero
seqtk
seqtools
seriation
setcover
sga
shasta
shovill
sibelia
sibsim4
sickle
sight
sigma-align
sigviewer
sim4
simde
simka
simrisc
ska
skesa
skewer
smalt
smrtanalysis
snakemake
snap
snap-aligner
sniffles
snippy
snpeff
snpomatic
snpsift
snp-sites
soapaligner
soapdenovo
soapdenovo2
soapsnp
socket++
sorted-nearest
sortmerna
spaced
spades
spaln
spdlog
spoa
sprai
spread-phy
sra-sdk
srf
srst2
ssake
sspace
ssshtest
stacks
staden
staden-io-lib
stringtie
subarch-select
subread
suitename
sumaclust
sumalibs
sumatra
surankco
surpyvor
survivor
svim
swarm-cluster
sweed
swissknife
tantan
tao-config
tao-json
t-coffee
terraphast
theseus
thesias
tiddit
tigr-glimmer
tipp
tm-align
tnseq-transit
tombo
toml11
tophat-recondition
toppred
tortoize
trace2dbest
tracetuner
transdecoder
transrate-tools
transtermhp
treeviewx
trf
trim-galore
trimmomatic
trinculo
trinityrnaseq
tvc
uc-echo
ugene
umis
uncalled
unifrac
unikmer
varna
vcfanno
vcftools
velvet
velvetoptimiser
veryfasttree
virulencefinder
vmatch
volpack
vsearch
vt
vtk-dicom
wham-align
wise
wtdbg2
xdffileio
xenium
xmedcon
xpore
xxsds-dynamic
yaggo
yaha
yanagiba
yanosim
912 (92.9%) packages
successfully build reproducibly:
abacas
abpoa
abyss
adapterremoval
adun.app
aegean
aeonbits-owner
aevol
aghermann
alien-hunter
allelecount
alter-sequence-alignment
altree
amap-align
amide
ampliconnoise
andi
ants
any2fasta
aragorn
arden
argh
argtable2
ariba
artemis
artfastqgenerator
art-nextgen-simulation-tools
assembly-stats
assemblytics
ataqv
atropos
augur
augustus
autodocksuite
autodock-vina
axe-demultiplexer
baitfisher
bali-phy
bambamc
bamclipper
bamkit
bamtools
bandage
barrnap
bart
bart-view
bbhash
bbmap
bcalm
bcftools
beads
beagle
beast-mcmc
bedops
bedtools
berkeley-express
bifrost
bioawk
biobambam2
bio-eagle
biojava4-live
biojava6-live
biojava-live
biomaj3
biomaj3-cli
biomaj3-core
biomaj3-daemon
biomaj3-download
biomaj3-process
biomaj3-user
biomaj3-zipkin
bioperl
bioperl-run
bio-rainbow
biosig
biosquid
biosyntax
bio-tradis
bio-vcf
bitseq
blasr
bolt-lmm
bowtie
bowtie2
boxshade
bppphyview
bppsuite
brian
brig
busco
bustools
bwa
camitk
camp
canu
capsule-maven-nextflow
capsule-nextflow
cassiopee
castxml
cat-bat
cct
cdbfasta
cde
cd-hit
centrifuge
cgview
changeo
charls
chip-seq
chromhmm
chromimpute
ciftilib
cif-tools
circlator
circos
circos-tools
civetweb
clearcut
clonalframe
clonalframeml
clonalorigin
clustalo
clustalw
clustalx
cnvkit
codonw
coils
concavity
concurrentqueue
conda-package-handling
conda-package-streaming
conservation-code
covtobed
crac
ctdconverter
ctdopts
ctffind
ctn
ctsim
cutesv
cwlformat
cwltest
cycle
cyvcf2
daligner
damapper
dascrubber
dawg
dazzdb
dcm2niix
dcmstack
debian-med
deblur
delly
density-fitness
dextractor
dialign
dialign-t
diamond-aligner
dicomscope
discosnp
disulfinder
dnaclust
dnapi
dnarrange
drmaa
drop-seq
dssp
dwgsim
ea-utils
ecopcr
edfbrowser
edflib
edtsurf
eegdev
eigensoft
elastix
elph
embassy-domainatrix
embassy-domalign
embassy-domsearch
emboss-explorer
e-mem
emmax
emperor
epcr
estscan
examl
exonerate
fast5
fasta3
fastani
fastaq
fastdnaml
fastlink
fastml
fastp
fastqc
fastq-pair
fastqtl
fasttree
fermi-lite
ffindex
figtree
fitgcp
flash
flexbar
flye
freecontact
fsa
fsm-lite
g2
galileo
garli
gasic
gatk-bwamem
gatk-fermilite
gdpc
gemma
genometester
genomethreader
genometools
genomicsdb
gentle
getdata
gfapy
gff2aplot
gff2ps
gffread
ggd-utils
ghmm
gifticlib
gjh-asl-json
glam2
gmap
gnumed-client
gnumed-server
grabix
graphlan
grinder
gsort
gubbins
gwama
gwyddion
harmonypy
harvest-tools
hdmf
hhsuite
hilive
hinge
hisat2
hmmer2
hnswlib
hopscotch-map
htscodecs
htseq
htsjdk
htslib
hts-nim-tools
hunspell-en-med
hyphy
icb-utils
idba
idseq-bench
igor
igv
iitii
illustrate
imagej
imbalanced-learn
indelible
infernal
insighttoolkit5
insilicoseq
invesalius
ipig
ismrmrd#
itkadaptivedenoising
itkgenericlabelinterpolator
iva
ivar
jaligner
jam-lib
jellyfish
jellyfish1
jheatchart
jmodeltest
kalign
kallisto
kaptive
kineticstools
king
king-probe
kissplice
klustakwik
kma
kmc
kmer
kmerresistance
kraken
kraken2
lagan
lamarc
lamassemble
lambda-align
lambda-align2
last-align
lastz
lefse
libace-perl
libamplsolver
libargparse
libargs
libatomicbitvector
libatomic-queue
libbigwig
libbio-alignio-stockholm-perl
libbio-cluster-perl
libbio-coordinate-perl
libbio-das-lite-perl
libbio-db-biofetch-perl
libbio-db-embl-perl
libbio-db-hts-perl
libbio-db-ncbihelper-perl
libbio-db-seqfeature-perl
libbio-featureio-perl
libbio-graphics-perl
libbio-mage-perl
libbio-mage-utils-perl
libbioparser-dev
libbiosoup-dev
libbio-tools-run-remoteblast-perl
libbio-variation-perl
libbitarray
libbpp-core
libbpp-phyl
libbpp-phyl-omics
libbpp-popgen
libbpp-qt
libbpp-raa
libbpp-seq
libbpp-seq-omics
libcereal
libchado-perl
libcolt-free-java
libctapimkt
libdisorder
libdistlib-java
libdivsufsort
libedlib
libfastahack
libflathashmap
libfreecontact-perl
libgclib
libgdf
libgenome
libgenome-model-tools-music-perl
libgenome-perl
libgff
libgkarrays
libgoby-java
libgo-perl
libgtextutils
libgzstream
libhac-java
libhat-trie
libhmsbeagle
libhpptools
libhttp-nio-java
libics
libips4o
libjloda-java
libjung-free-java
libkmlframework-java
libla4j-java
libleidenalg
libmaus2
libmcfp
libmems
libmialm
libminc
libmjson-java
libmmap-allocator
libmmmulti
libmurmurhash
libmuscle
libncl
libnewuoa
libomp-jonathonl
liboptions-java
libpal-java
libpdb-redo
libpj-java
libpsortb
libqes
librandom123
librcsb-core-wrapper
librdp-taxonomy-tree-java
librg-blast-parser-perl
librg-exception-perl
librg-utils-perl
librostlab-blast
libsecrecy
libseqlib
libshrinkwrap
libsis-base-java
libslow5lib
libsmithwaterman
libsort-key-top-perl
libssw
libstatgen
libstreamvbyte
libtabixpp
libtecla
libtfbs-perl
libthread-pool
libvbz-hdf-plugin
libvcflib
libvistaio
libwfa2
libxdf
libzeep
libzerg
libzerg-perl
lighter
loki
ltrsift
lucy
lumpy-sv
macs
maffilter
mafft
malt
mapdamage
maq
maqview
mash
maude
mauve-aligner
maxflow
mcaller
mcl
mcl14
mecat2
medicalterms
megadepth
megahit
megan-ce
melting
mencal
metabat
metaeuk
metaphlan
metaphlan2-data
metastudent
metastudent-data-2
mhap
mialmpick
microbegps
microbiomeutil
milib
minc-tools
mindthegap
minia
miniasm
minimac4
minimap
minimap2
mipe
mira
mirtop
mmlib
mmseqs2
mosdepth
mothur
mptp
mrbayes
mricron
multiqc
mummer
murasaki
muscle
muscle3
mustang
nanofilt
nanolyse
nanook
nanopolish
nanostat
nanosv
ncbi-acc-download
ncbi-entrez-direct
ncbi-seg
ncbi-tools6
ncbi-vdb
neo
neobio
ngmlr
nibabel
nifticlib
nim-hts
nim-kexpr
nim-lapper
nipy
nipype
njplot
norsnet
norsp
ntcard
nthash
nutsqlite
nxtrim
obitools
odin
ont-fast5-api
opencfu
openslide
openslide-python
optimir
orthanc
orthanc-dicomweb
orthanc-gdcm
orthanc-imagej
orthanc-mysql
orthanc-neuro
orthanc-postgresql
orthanc-python
orthanc-webviewer
orthanc-wsi
oscar
pairtools
pal2nal
paleomix
paml
papyrus
paraclu
parafly
parallel-fastq-dump
parasail
parsinsert
parsnp
paryfor
patman
patsy
pbbam
pbcopper
pbdagcon
pbseqlib
pbsim
pbsuite
pcalendar
pdb2pqr
peptidebuilder
perlprimer
perm
pftools
phast
phipack
phybin
phylip
phylonium
phyml
physamp
phyutility
phyx
picard-tools
picopore
pigx-rnaseq
piler
pilercr
pilon
pinfish
pique
pirs
pixelmed
pixelmed-codec
pizzly
placnet
plasmidid
plasmidomics
plasmidseeker
plast
plastimatch
plink
plink1.9
plink2
plip
poa
populations
poretools
praat
prank
predictnls
presto
prime-phylo
primer3
prinseq-lite
proalign
probabel
probalign
probcons
proda
prodigal
profbval
profisis
profnet
profphd-utils
proftmb
progressivemauve
prokka
propka
proteinortho
prottest
provean
pscan-chip
pscan-tfbs
psignifit
psortb
pullseq
pvrg-jpeg
pybel
pybigwig
pychopper
pycoqc
pycorrfit
pyensembl
pyfastx
pynwb
pyode
pyqi
pyscanfcs
python-alignlib
python-anndata
python-awkward
python-bcbio-gff
python-bel-resources
python-bids-validator
python-bioblend
python-bioframe
python-biom-format
python-biopython
python-bioregistry
python-biotools
python-bx
python-cgecore
python-cgelib
python-cigar
python-cobra
python-cooler
python-csb
python-curies
python-cutadapt
python-cykhash
python-datacache
python-deeptools
python-deeptoolsintervals
python-dendropy
python-dicompylercore
python-dnaio
python-epimodels
python-ete3
python-etelemetry
python-fitbit
python-freecontact
python-geneimpacts
python-gffutils
python-gtfparse
python-hl7
python-hmmlearn
python-intervaltree-bio
python-iow
python-leidenalg
python-mne
python-nanoget
python-nanomath
python-pairix
python-pangolearn
python-parasail
python-pauvre
python-pbcommand
python-pbcore+
python-prefixed
python-py2bit
python-pyani
python-pybedtools
python-pycosat
python-pyfaidx
python-pymummer
python-pyspoa
python-pyvcf
python-questplus
python-ruffus
python-scitrack
python-screed
python-skbio
python-sqt
python-tinyalign
python-treetime
python-trx-python
python-wdlparse
pyxid
pyxnat
q2-alignment
q2cli
q2-cutadapt
q2-dada2
q2-demux
q2-diversity-lib
q2-emperor
q2-feature-table
q2-fragment-insertion
q2-metadata
q2-phylogeny
q2-quality-control
q2-quality-filter
q2-sample-classifier
q2-taxa
q2templates
q2-types
qcat
qcumber
qiime
qrisk2
qsopt-ex
qtltools
quicktree
quitcount
quorum
racon
radiant
ragout
rambo-k
rampler
rapmap
raster3d
rate4site
raxml
ray
rdp-alignment
rdp-classifier
rdp-readseq
readerwriterqueue
readseq
readucks
reapr
recan
relacy
relion
repeatmasker-recon
reprof
resfinder
resfinder-db
rnahybrid
rna-star
roary
rockhopper
roguenarok
routine-update
rsem
rtax
ruby-rgfa
runcircos-gui
saint
salmid
sambamba
samblaster
samclip
samtools
samtools-legacy
savvy
sbmltoolbox
scoary
scrappie
scrm
scythe
seaview
seer
seirsplus
sepp
seqan-needle
seqan-raptor
seqkit
seqmagick
seqprep
seqsero
seqtk
seqtools
seriation
setcover
sga
shasta
shovill
sibelia
sibsim4
sickle
sight
sigma-align
sigviewer
sim4
simde
simka
simrisc
ska
skesa
skewer
smalt
smrtanalysis
snakemake
snap
snap-aligner
sniffles
snippy
snpeff
snpomatic
snpsift
snp-sites
soapaligner
soapdenovo
soapdenovo2
soapsnp
socket++
sorted-nearest
sortmerna
spaced
spades
spaln
spdlog
spoa
sprai
spread-phy
sra-sdk
srf
srst2
ssake
sspace
ssshtest
stacks
staden
staden-io-lib
stringtie
subarch-select
subread
suitename
sumaclust
sumalibs
sumatra
surankco
surpyvor
survivor
svim
swarm-cluster
sweed
swissknife
tantan
tao-config
tao-json
t-coffee
terraphast
theseus
thesias
tiddit
tigr-glimmer
tipp
tm-align
tnseq-transit
tombo
toml11
tophat-recondition
toppred
tortoize
trace2dbest
tracetuner
transdecoder
transrate-tools
transtermhp
treeviewx
trf
trim-galore
trimmomatic
trinculo
trinityrnaseq
tvc
uc-echo
ugene
umis
uncalled
unifrac
unikmer
varna
vcfanno
vcftools
velvet
velvetoptimiser
veryfasttree
virulencefinder
vmatch
volpack
vsearch
vt
vtk-dicom
wham-align
wise
wtdbg2
xdffileio
xenium
xmedcon
xpore
xxsds-dynamic
yaggo
yaha
yanagiba
yanosim
A package name displayed with a bold
font is an indication that this package has a note. Visited
packages are linked in green, those which have not been visited are
linked in blue.
A # sign after the name of a package
indicates that a bug is filed against it. Likewise, a
+ sign indicates there is a
patch available, a P means a
pending bug while # indicates a
closed bug. In cases of several bugs, the symbol is repeated.