maint_debian-med package set for buster/amd64
Debian package sets:
desktop package sets:
Debian distribution package sets:
maintenance team package sets:
-
 maint_debian-accessibility
maint_debian-accessibility
-
 maint_debian-boot
maint_debian-boot
-
 maint_debian-lua
maint_debian-lua
-
 maint_debian-med
maint_debian-med
-
 maint_debian-ocaml
maint_debian-ocaml
- maint_debian-on-mobile-maintainers
-
 maint_debian-python
maint_debian-python
-
 maint_debian-qa
maint_debian-qa
-
 maint_debian-science
maint_debian-science
-
 maint_debian-x
maint_debian-x
-
 maint_pkg-android-tools-devel
maint_pkg-android-tools-devel
-
 maint_pkg-fonts-devel
maint_pkg-fonts-devel
-
 maint_pkg-games-devel
maint_pkg-games-devel
-
 maint_pkg-golang-maintainers
maint_pkg-golang-maintainers
-
 maint_pkg-grass-devel
maint_pkg-grass-devel
-
 maint_pkg-haskell-maintainers
maint_pkg-haskell-maintainers
-
 maint_pkg-java-maintainers
maint_pkg-java-maintainers
-
 maint_pkg-javascript-devel
maint_pkg-javascript-devel
-
 maint_pkg-multimedia-maintainers
maint_pkg-multimedia-maintainers
-
 maint_pkg-perl-maintainers
maint_pkg-perl-maintainers
-
 maint_pkg-php-pear
maint_pkg-php-pear
-
 maint_pkg-openstack
maint_pkg-openstack
- maint_pkg-r
-
 maint_pkg-ruby-extras-maintainers
maint_pkg-ruby-extras-maintainers
- maint_pkg-rust-maintainers
- maint_reproducible-builds
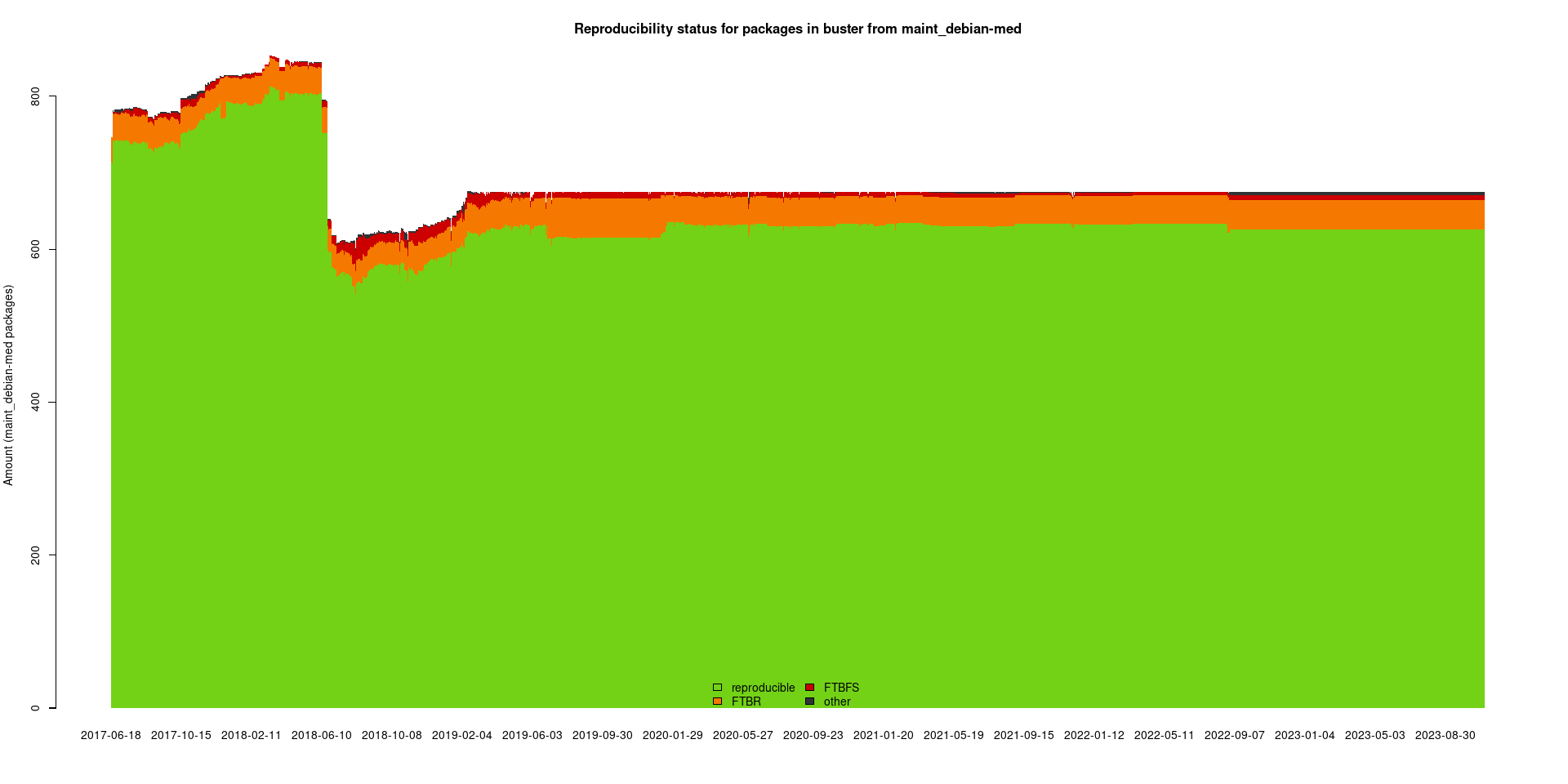
The package set maint_debian-med in
buster/amd64 consists of 675 packages:
 38 (5.6%) packages
failed to build reproducibly:
sleepyhead
emboss
treeview
camitk
flexbar
mapsembler2
lambda-align
macsyfinder
dicom3tools
fastx-toolkit
mcl
librostlab
iqtree
mira
bowtie2
ffindex
metastudent-data
microbiomeutil
mummer
centrifuge
jmodeltest
pixelmed
ncbi-blast+
htslib
crac
fis-gtm
gbrowse+
gdcm
grabix
logol
libsbml
raster3d
qtltools
sra-sdk
trinityrnaseq
toil
gatb-core
ball
38 (5.6%) packages
failed to build reproducibly:
sleepyhead
emboss
treeview
camitk
flexbar
mapsembler2
lambda-align
macsyfinder
dicom3tools
fastx-toolkit
mcl
librostlab
iqtree
mira
bowtie2
ffindex
metastudent-data
microbiomeutil
mummer
centrifuge
jmodeltest
pixelmed
ncbi-blast+
htslib
crac
fis-gtm
gbrowse+
gdcm
grabix
logol
libsbml
raster3d
qtltools
sra-sdk
trinityrnaseq
toil
gatb-core
ball
 7 (1.0%) packages
failed to build from source:
python-biopython
seqan
seqan2
libmurmurhash
jellyfish
simpleitk
gemma
7 (1.0%) packages
failed to build from source:
python-biopython
seqan
seqan2
libmurmurhash
jellyfish
simpleitk
gemma



 4 (0.6%) packages
are either in depwait state, blacklisted, not for us, or cannot be downloaded:
python-pysam
pbbam
ismrmrd#
ngs-sdk
4 (0.6%) packages
are either in depwait state, blacklisted, not for us, or cannot be downloaded:
python-pysam
pbbam
ismrmrd#
ngs-sdk
 626 (92.7%) packages
successfully build reproducibly:
abacas
abyss
acedb
adapterremoval
adun.app
aegean
aeskulap
aevol
aghermann
alien-hunter
alter-sequence-alignment
altree
amap-align
amide
ampliconnoise
andi
anfo
aragorn
arden
ariba
artemis
artfastqgenerator
art-nextgen-simulation-tools
assemblytics
augustus
autodocksuite
autodock-vina
axe-demultiplexer
baitfisher
bali-phy
bambamc
bamtools
bandage
barrnap
bart
bart-view
bcbio
bcftools
beads
beagle
beast2-mcmc
beast-mcmc
bedops
bedtools
berkeley-express
biococoa
bio-eagle
biojava4-live
biojava-live
biomaj3
biomaj3-cli
biomaj3-core
biomaj3-daemon
biomaj3-download
biomaj3-process
biomaj3-user
biomaj3-zipkin
bioperl
bioperl-run
bio-rainbow
biosquid
biosyntax
bio-tradis
bitseq
blasr
bolt-lmm
bowtie
boxshade
bppphyview
bppsuite
brig
bwa
cain
camp
canu
cassiopee
cct
cdbfasta
cd-hit
cfflib
cgview
changeo
charls
chromhmm
chromimpute
ciftilib
circlator
circos
circos-tools
clearcut
clonalframe
clonalframeml
clonalorigin
clustalo
clustalw
clustalx
cnvkit
codonw
coils
concavity
consensuscore
conservation-code
ctdconverter
ctdopts
ctn
ctsim
cwltool
cycle
cyvcf2
daligner
dascrubber
dawg
dazzdb
dcm2niix
dcmtk
debian-med
deepnano
delly
dialign
dialign-t
diamond-aligner
dicompyler
dicomscope
dindel
discosnp
disulfinder
dnaclust
dssp
dwgsim
ea-utils
ecopcr
edfbrowser
edflib
edtsurf
eigensoft
elastix
elph
embassy-domainatrix
embassy-domalign
embassy-domsearch
emboss-explorer
e-mem
epcr
estscan
examl
exonerate
fast5
fastaq
fastdnaml
fastlink
fastml
fastp
fastqc
fastqtl
fasttree
fermi-lite
figtree
fitgcp
freebayes
freecontact
freemedforms-project
fsa
fsm-lite
fw4spl
g2
galileo
garli
gasic
gdpc
genometester
genometools
gentle
getdata
gfapy
gff2aplot
gff2ps
ghmm
giira
ginkgocadx
glam2
gnumed-client
gnumed-server
golang-github-dataence-porter2
graphlan
grinder
gubbins
gwama
gwyddion
harvest-tools
hhsuite
hilive
hinge
hisat2
hmmer
hmmer2
htseq
htsjdk
hunspell-en-med
hyphy
idba
igor
igraph
imagej
imagevis3d
indelible
infernal
insighttoolkit4
invesalius
ipig
iva
jaligner
jam-lib
jebl2
jellyfish1
jheatchart
kalign
khmer
kineticstools
king
king-probe
kissplice
kmc
kmer
kraken
lagan
lamarc
lambda-align2
last-align
lefse
libace-perl
libbio-coordinate-perl
libbio-das-lite-perl
libbio-graphics-perl
libbio-mage-perl
libbio-mage-utils-perl
libbioparser-dev
libbpp-core
libbpp-phyl
libbpp-phyl-omics
libbpp-popgen
libbpp-qt
libbpp-raa
libbpp-seq
libbpp-seq-omics
libcereal
libchado-perl
libcolt-free-java
libctapimkt
libdeflate
libdisorder
libdivsufsort
libedlib
libfastahack
libfreecontact-perl
libgenome
libgenome-model-tools-music-perl
libgenome-perl
libgff
libgkarrays
libgo-perl
libgtextutils
libgzstream
libhac-java
libhat-trie
libhmsbeagle
libhpptools
libics
libjbzip2-java
libjloda-java
libjung-free-java
libkmlframework-java
liblemon
libmems
libmialm
libminc
libmuscle
libncl
liboptions-java
libpal-java
libpll
libpsortb
libqes
libquazip
librandom123
librcsb-core-wrapper
librdp-taxonomy-tree-java
librg-blast-parser-perl
librg-exception-perl
librg-utils-perl
librostlab-blast
libseqlib
libsis-base-java
libsmithwaterman
libsort-key-top-perl
libssw
libstatgen
libtabixpp
libtecla
libtfbs-perl
libthread-pool
libundead
libvcflib
libvistaio
libxdf
libzeep
libzerg
libzerg-perl
libzstd
loki
ltrsift
lucy
macs
maffilter
mafft
mapdamage
maq
maqview
mash
maude
mauve-aligner
maxflow
medicalterms
melting
mencal
metaphlan2
metaphlan2-data
metastudent
metastudent-data-2
mhap
mia
mialmpick
miaviewit
microbegps
milib
minc-tools
minia
miniasm
minimac4
minimap
minimap2
mipe
mlv-smile
mobyle
mobyle-programs
mobyle-tutorials
mothur
mptp
mrbayes
mriconvert
mrs
murasaki
muscle
mustang
mypy
nanook
nanopolish
ncbi-entrez-direct
ncbi-seg
ncbi-tools6
ncbi-vdb
neobio
njplot
norsnet
norsp
nutsqlite
obitools
odil
opencfu
openslide
openslide-python
opensurgsim
orthanc
orthanc-dicomweb
orthanc-imagej
orthanc-mysql
orthanc-postgresql
orthanc-webviewer
orthanc-wsi
pal2nal
paleomix
paml
papyrus
paraclu
parafly
parsinsert
parsnp
patman
pbalign
pbbarcode
pbcopper
pbdagcon
pbgenomicconsensus
pbh5tools
pbseqlib
pbsim
pbsuite
pcalendar
pdb2pqr
perlprimer
perm
pftools
phast
phipack
phybin
phylip
phyml
physamp
phyutility
phyx
picard-tools
piler
pilon
pirs
pixelmed-codec
placnet
plasmidomics
plasmidseeker
plast
plastimatch
plink
plink1.9
plip
poa
pondus
populations
porechop
poretools
pp-popularity-contest
praat
prank
predictnls
predictprotein
prime-phylo
primer3
proalign
probabel
probalign
probcons
proda
prodigal
profbval
profisis
profnet
profphd
profphd-utils
proftmb
progressivemauve
proteinortho
prottest
pscan-chip
pscan-tfbs
psortb
pvrg-jpeg
pycorrfit
pymia
pynast
pyqi
pyscanfcs
python3-typed-ast
python-avro
python-biom-format
python-biotools
python-burrito
python-bx
python-bz2file
python-clips
python-cobra
python-cogent
python-colormap
python-colormath
python-csb
python-cutadapt
python-dendropy
python-depinfo
python-easydev
python-fitbit
python-freecontact
python-gffutils
python-hdmedians
python-hl7
python-intervaltree-bio
python-lzstring
python-matplotlib-venn
python-mne
python-multipletau
python-mypy-extensions
python-pbcommand
python-pbcore
python-pipdeptree
python-pybedtools
python-pyfaidx
python-pyflow
python-pymummer
python-pyvcf
python-qcli
python-rdflib-jsonld
python-ruffus
python-schema-salad
python-screed
python-spectra
python-sqlsoup
python-sqt
python-treetime
python-xopen
qcumber
qrisk2
qsopt-ex
quorum
racon
radiant
rambo-k
rampler
rapmap
rate4site
raxml
ray
rdp-alignment
rdp-classifier
rdp-readseq
readseq
reapr
relion
repeatmasker-recon
reprof
rnahybrid
rna-star
roary
roguenarok
rsem
rtax
ruby-rgfa
runcircos-gui
saint
salmid
salmon
samblaster
samtools
samtools-legacy
sbmltoolbox
scoary
scrm
scythe
seaview
seer
segemehl
seqmagick
seqprep
seqsero
seqtk
seqtools
sga
sibsim4
sickle
sigma-align
sim4
smalr
smalt
snakemake
snap
snap-aligner
sniffles
snpomatic
snp-sites
soapaligner
soapdenovo
soapdenovo2
soapsnp
socket++
sofa-framework
sortmerna
spaced
spades
spdlog
sphinxcontrib-autoprogram
spoa
sprai
spread-phy
squizz
srf
srst2
ssake
sspace
stacks
staden
staden-io-lib
subread
suitename
sumaclust
sumatra
surankco
swarm-cluster
sweed
swissknife
tantan
t-coffee
theseus
tifffile
tigr-glimmer
tm-align
tnseq-transit
tophat
toppred
trace2dbest
tracetuner
transdecoder
transrate-tools
transtermhp
tree-puzzle
treeviewx
trimmomatic
tvc
uc-echo
unanimity
unicycler
varna
vcftools
velvet
velvetoptimiser
volpack
vsearch
vtk-dicom
wise
xmedcon
yaggo
yaha
zalign
626 (92.7%) packages
successfully build reproducibly:
abacas
abyss
acedb
adapterremoval
adun.app
aegean
aeskulap
aevol
aghermann
alien-hunter
alter-sequence-alignment
altree
amap-align
amide
ampliconnoise
andi
anfo
aragorn
arden
ariba
artemis
artfastqgenerator
art-nextgen-simulation-tools
assemblytics
augustus
autodocksuite
autodock-vina
axe-demultiplexer
baitfisher
bali-phy
bambamc
bamtools
bandage
barrnap
bart
bart-view
bcbio
bcftools
beads
beagle
beast2-mcmc
beast-mcmc
bedops
bedtools
berkeley-express
biococoa
bio-eagle
biojava4-live
biojava-live
biomaj3
biomaj3-cli
biomaj3-core
biomaj3-daemon
biomaj3-download
biomaj3-process
biomaj3-user
biomaj3-zipkin
bioperl
bioperl-run
bio-rainbow
biosquid
biosyntax
bio-tradis
bitseq
blasr
bolt-lmm
bowtie
boxshade
bppphyview
bppsuite
brig
bwa
cain
camp
canu
cassiopee
cct
cdbfasta
cd-hit
cfflib
cgview
changeo
charls
chromhmm
chromimpute
ciftilib
circlator
circos
circos-tools
clearcut
clonalframe
clonalframeml
clonalorigin
clustalo
clustalw
clustalx
cnvkit
codonw
coils
concavity
consensuscore
conservation-code
ctdconverter
ctdopts
ctn
ctsim
cwltool
cycle
cyvcf2
daligner
dascrubber
dawg
dazzdb
dcm2niix
dcmtk
debian-med
deepnano
delly
dialign
dialign-t
diamond-aligner
dicompyler
dicomscope
dindel
discosnp
disulfinder
dnaclust
dssp
dwgsim
ea-utils
ecopcr
edfbrowser
edflib
edtsurf
eigensoft
elastix
elph
embassy-domainatrix
embassy-domalign
embassy-domsearch
emboss-explorer
e-mem
epcr
estscan
examl
exonerate
fast5
fastaq
fastdnaml
fastlink
fastml
fastp
fastqc
fastqtl
fasttree
fermi-lite
figtree
fitgcp
freebayes
freecontact
freemedforms-project
fsa
fsm-lite
fw4spl
g2
galileo
garli
gasic
gdpc
genometester
genometools
gentle
getdata
gfapy
gff2aplot
gff2ps
ghmm
giira
ginkgocadx
glam2
gnumed-client
gnumed-server
golang-github-dataence-porter2
graphlan
grinder
gubbins
gwama
gwyddion
harvest-tools
hhsuite
hilive
hinge
hisat2
hmmer
hmmer2
htseq
htsjdk
hunspell-en-med
hyphy
idba
igor
igraph
imagej
imagevis3d
indelible
infernal
insighttoolkit4
invesalius
ipig
iva
jaligner
jam-lib
jebl2
jellyfish1
jheatchart
kalign
khmer
kineticstools
king
king-probe
kissplice
kmc
kmer
kraken
lagan
lamarc
lambda-align2
last-align
lefse
libace-perl
libbio-coordinate-perl
libbio-das-lite-perl
libbio-graphics-perl
libbio-mage-perl
libbio-mage-utils-perl
libbioparser-dev
libbpp-core
libbpp-phyl
libbpp-phyl-omics
libbpp-popgen
libbpp-qt
libbpp-raa
libbpp-seq
libbpp-seq-omics
libcereal
libchado-perl
libcolt-free-java
libctapimkt
libdeflate
libdisorder
libdivsufsort
libedlib
libfastahack
libfreecontact-perl
libgenome
libgenome-model-tools-music-perl
libgenome-perl
libgff
libgkarrays
libgo-perl
libgtextutils
libgzstream
libhac-java
libhat-trie
libhmsbeagle
libhpptools
libics
libjbzip2-java
libjloda-java
libjung-free-java
libkmlframework-java
liblemon
libmems
libmialm
libminc
libmuscle
libncl
liboptions-java
libpal-java
libpll
libpsortb
libqes
libquazip
librandom123
librcsb-core-wrapper
librdp-taxonomy-tree-java
librg-blast-parser-perl
librg-exception-perl
librg-utils-perl
librostlab-blast
libseqlib
libsis-base-java
libsmithwaterman
libsort-key-top-perl
libssw
libstatgen
libtabixpp
libtecla
libtfbs-perl
libthread-pool
libundead
libvcflib
libvistaio
libxdf
libzeep
libzerg
libzerg-perl
libzstd
loki
ltrsift
lucy
macs
maffilter
mafft
mapdamage
maq
maqview
mash
maude
mauve-aligner
maxflow
medicalterms
melting
mencal
metaphlan2
metaphlan2-data
metastudent
metastudent-data-2
mhap
mia
mialmpick
miaviewit
microbegps
milib
minc-tools
minia
miniasm
minimac4
minimap
minimap2
mipe
mlv-smile
mobyle
mobyle-programs
mobyle-tutorials
mothur
mptp
mrbayes
mriconvert
mrs
murasaki
muscle
mustang
mypy
nanook
nanopolish
ncbi-entrez-direct
ncbi-seg
ncbi-tools6
ncbi-vdb
neobio
njplot
norsnet
norsp
nutsqlite
obitools
odil
opencfu
openslide
openslide-python
opensurgsim
orthanc
orthanc-dicomweb
orthanc-imagej
orthanc-mysql
orthanc-postgresql
orthanc-webviewer
orthanc-wsi
pal2nal
paleomix
paml
papyrus
paraclu
parafly
parsinsert
parsnp
patman
pbalign
pbbarcode
pbcopper
pbdagcon
pbgenomicconsensus
pbh5tools
pbseqlib
pbsim
pbsuite
pcalendar
pdb2pqr
perlprimer
perm
pftools
phast
phipack
phybin
phylip
phyml
physamp
phyutility
phyx
picard-tools
piler
pilon
pirs
pixelmed-codec
placnet
plasmidomics
plasmidseeker
plast
plastimatch
plink
plink1.9
plip
poa
pondus
populations
porechop
poretools
pp-popularity-contest
praat
prank
predictnls
predictprotein
prime-phylo
primer3
proalign
probabel
probalign
probcons
proda
prodigal
profbval
profisis
profnet
profphd
profphd-utils
proftmb
progressivemauve
proteinortho
prottest
pscan-chip
pscan-tfbs
psortb
pvrg-jpeg
pycorrfit
pymia
pynast
pyqi
pyscanfcs
python3-typed-ast
python-avro
python-biom-format
python-biotools
python-burrito
python-bx
python-bz2file
python-clips
python-cobra
python-cogent
python-colormap
python-colormath
python-csb
python-cutadapt
python-dendropy
python-depinfo
python-easydev
python-fitbit
python-freecontact
python-gffutils
python-hdmedians
python-hl7
python-intervaltree-bio
python-lzstring
python-matplotlib-venn
python-mne
python-multipletau
python-mypy-extensions
python-pbcommand
python-pbcore
python-pipdeptree
python-pybedtools
python-pyfaidx
python-pyflow
python-pymummer
python-pyvcf
python-qcli
python-rdflib-jsonld
python-ruffus
python-schema-salad
python-screed
python-spectra
python-sqlsoup
python-sqt
python-treetime
python-xopen
qcumber
qrisk2
qsopt-ex
quorum
racon
radiant
rambo-k
rampler
rapmap
rate4site
raxml
ray
rdp-alignment
rdp-classifier
rdp-readseq
readseq
reapr
relion
repeatmasker-recon
reprof
rnahybrid
rna-star
roary
roguenarok
rsem
rtax
ruby-rgfa
runcircos-gui
saint
salmid
salmon
samblaster
samtools
samtools-legacy
sbmltoolbox
scoary
scrm
scythe
seaview
seer
segemehl
seqmagick
seqprep
seqsero
seqtk
seqtools
sga
sibsim4
sickle
sigma-align
sim4
smalr
smalt
snakemake
snap
snap-aligner
sniffles
snpomatic
snp-sites
soapaligner
soapdenovo
soapdenovo2
soapsnp
socket++
sofa-framework
sortmerna
spaced
spades
spdlog
sphinxcontrib-autoprogram
spoa
sprai
spread-phy
squizz
srf
srst2
ssake
sspace
stacks
staden
staden-io-lib
subread
suitename
sumaclust
sumatra
surankco
swarm-cluster
sweed
swissknife
tantan
t-coffee
theseus
tifffile
tigr-glimmer
tm-align
tnseq-transit
tophat
toppred
trace2dbest
tracetuner
transdecoder
transrate-tools
transtermhp
tree-puzzle
treeviewx
trimmomatic
tvc
uc-echo
unanimity
unicycler
varna
vcftools
velvet
velvetoptimiser
volpack
vsearch
vtk-dicom
wise
xmedcon
yaggo
yaha
zalign
A package name displayed with a bold
font is an indication that this package has a note. Visited
packages are linked in green, those which have not been visited are
linked in blue.
A # sign after the name of a package
indicates that a bug is filed against it. Likewise, a
+ sign indicates there is a
patch available, a P means a
pending bug while # indicates a
closed bug. In cases of several bugs, the symbol is repeated.